SARS-CoV-2
21-03-2022
Unheeded SARS-CoV-2 proteins? A deep look into negative-sense RNA
Our new research is just out in the Briefings in Bioinformatics!
https://doi.org/10.1093/bib/bbac045
![Localization and synteny of nsORFs in SARS-CoV-2 and related coronaviruses. (A) Localization of all identified nsORFs within the SARS-CoV-2 genome. The upper part of the scheme depicts positively encoded ORFs annotated on NCBI reference SARS-CoV-2 genome, together with additional ORFs described in the literature (indicated in italics): ORF3c, ORF3d, ORF3b (which span particular genomic regions of ORF3a) and ORF9b and ORF9c (which span particular genomic regions of N). (B)Synteny of SARS-CoV-2 nsORFs in representative species of SARS-like coronaviruses. At least two SARS-CoV-2 nsORFs (nsORF2 and nsORF9) are more or less conserved in most of inspected SARS-CoV-2-related coronaviruses, including old SARS-CoV Tor 2003. In MERS-CoV and human-CoV-OC43, none of SARS-CoV-2 homologous nsORFs was found. The synteny plot was constructed using SimpleSynteny web server [56] and redrawn in this schematic figure.](https://oup.silverchair-cdn.com/oup/backfile/Content_public/Journal/bib/PAP/10.1093_bib_bbac045/2/bbac045f1.jpeg?Expires=1650881874&Signature=N28~yPS2hlDh7UjhnwKhx70g4MjwwwRVprE8Mc32hwxnbAwnuDvrAudlEnKomLu88Pyt9ox2YzCkGpdYQ83A3rScUP~1laJoh9T2Kcst8xMyk5lz07PlE9Dp0REVvima5jlugybcfH2V1rLTDj~vT3lp1Eq~T02yeuRdn4Vl2z4y1rluSIrascW0VNWZNMswFXEFGU~nkSAMpaq~QHUEVBW~bIB-oeF~eR0j9GixB1NrrpB28qweoL6EUnWK92gKB1Ku2pIrcyZcFjDsMleHgiN9Aqn2OzE3-Lw3nPJBlqOs7CaUgQjIsPJiV2xiRUKgjvUxvPwCPG5st1plOr6eJQ__&Key-Pair-Id=APKAIE5G5CRDK6RD3PGA)
09-04-2021
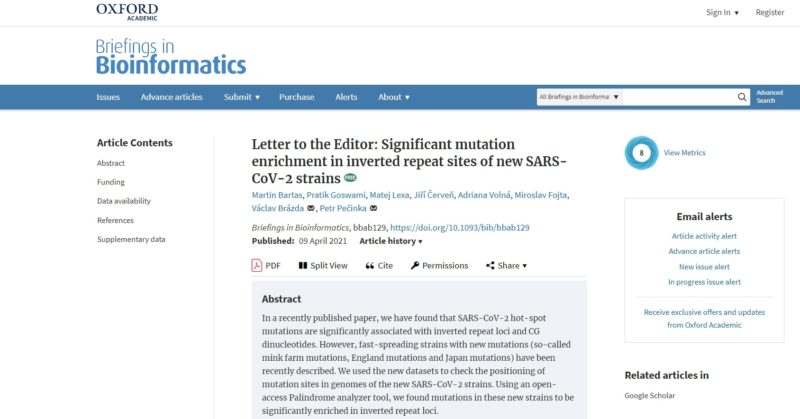
Follow-up study just published!!!
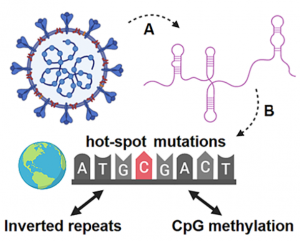
Inverted repeats in coronaviral RNA are a rich source of SARS-CoV-2 genetic instability. It was found that novel SARS-CoV-2 strains (England, Brazil,...) characteristic mutations are also overrepresented within inverted repeats sites. This can be utilized for the prediction of future mutation occurrence.
Read for free (open-access):
https://academic.oup.com/bib/advance-article/doi/10.1093/bib/bbab129/6219140
21-12-2020
Our team is intensively working in the field of novel coronavirus research.
SARS-CoV-2 is an intensively investigated virus from the order Nidovirales (Coronaviridae family) that causes COVID-19 disease in humans. Through enormous scientific effort, thousands of viral strains have been sequenced to date, thereby creating a strong background for deep bioinformatics studies of the SARS-CoV-2 genome. In our recent study published in the Briefings in Bioinformatics journal (https://academic.oup.com/bib/advance-article/doi/10.1093/bib/bbaa385/6042389), we have found that SARS-CoV-2 hot-spot mutations are significantly enriched within both inverted repeats and CpG island loci. This points to their role in genomic instability and may predict further mutational drive of the SARS-CoV-2 genome. Moreover, SARS-CoV-2 CpG islands are strongly enriched upstream from viral ORFs and thus could play important roles in transcription and the viral life cycle. We hypothesize that hypermethylation of these loci will decrease the transcription of viral ORFs and could therefore limit the progression of the disease.